The cytoverse is a tightly coordinated collection of R packages striving to make cytometry data analysis easier and more computationally efficient.
flowWorkspace
flowWorkspace provides core data structures and methods for use in cytometry data analysis, including compensation, transformation, and gating.
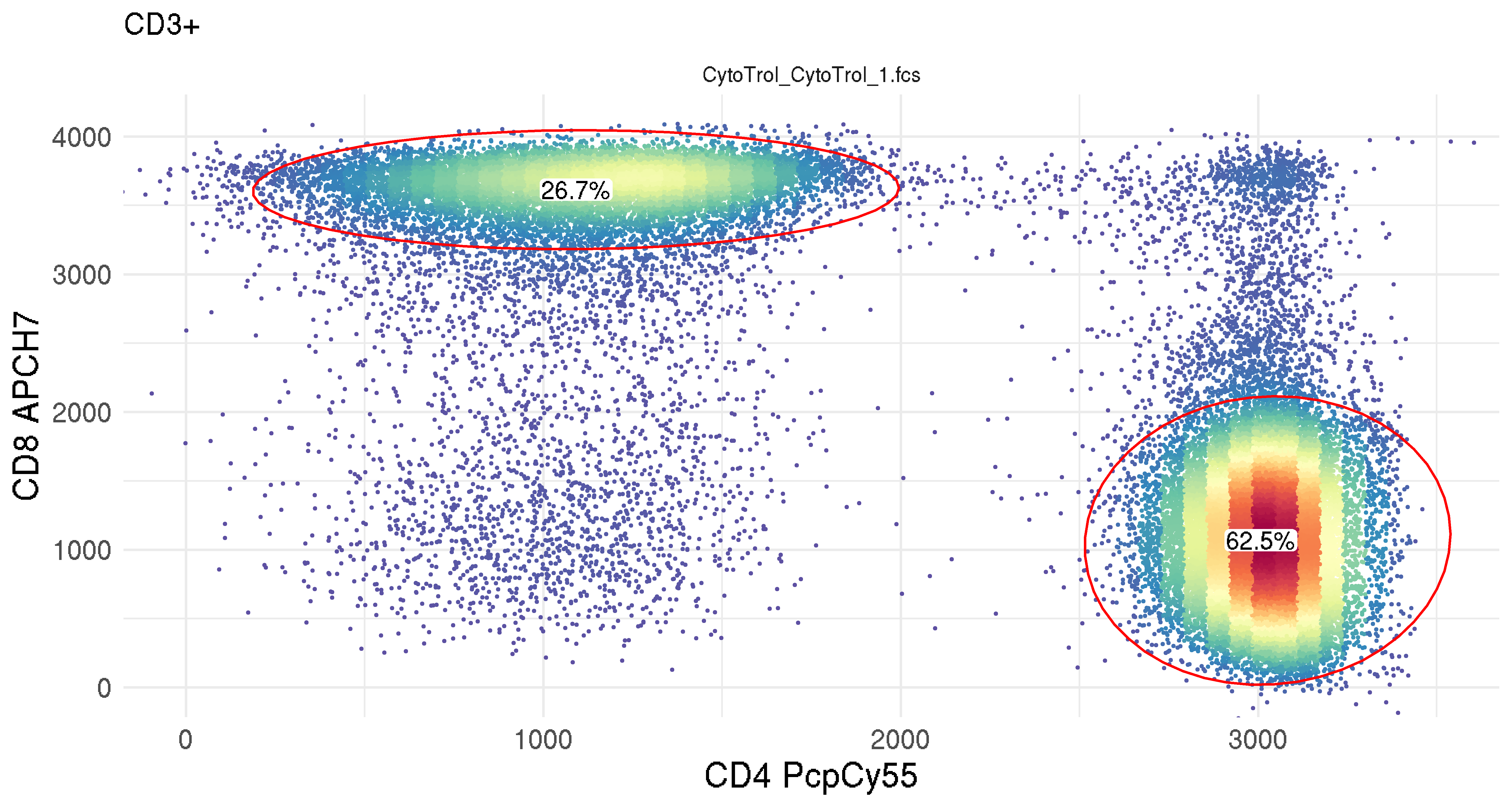
ggcyto
ggcyto provides cytometry-specific plotting functionality mirroring ggplot2 semantics.
CytoML
CytoML allows for import and export of gating analyses of cytometry data to or from FlowJo, BD FACSDIVA, or Cytobank workspace formats.
> ws <- open_diva_xml(ws_file)
> gs <- diva_to_gatingset(ws)
...
> gatingset_to_flowjo(gs, flowjo_filename)
openCyto
openCyto enables the development of reproducible automated analysis pipelines.
A number of packages either directly support or extend the core cytoverse packages.
- RProtoBufLib
- cytolib
- flowCore